Overview
Teaching: 20 min
Exercises: 25 minQuestions
Can I work with data from multiple sources?
How can I combine data from different data sets?
Objectives
Combine data from multiple files into a single DataFrame using merge and concat.
Combine two DataFrames using a unique ID found in both DataFrames.
Employ
to_csvto export a DataFrame in CSV format.Join DataFrames using common fields (join keys).
In many “real world” situations, the data that we want to use come in multiple
files. We often need to combine these files into a single DataFrame to analyze
the data. The pandas package provides various methods for combining
DataFrames including
merge and concat.
To work through the examples below, we first need to load the species and surveys files into pandas DataFrames. In iPython:
import pandas as pd
sep_data = pd.read_csv("pore_info.tsv", delimiter='\t')
sep_data['Run'] = "Not great run"
[395 rows x 12 columns]
good_data = pd.read_csv("good_data.tsv",
delimiter='\t')
good_data['Run'] = "Great run"
[395 rows x 12 columns]
Concatenating DataFrames
We can use the concat function in Pandas to append either columns or rows from
one DataFrame to another. Let’s grab two subsets of our data to see how this
works.
To stack the data vertically, we need to make sure we have the
same columns and associated column format in both datasets. When we stack
horizonally, we want to make sure what we are doing makes sense (ie the data are
related in some way).
# Stack the DataFrames on top of each other
vertical_stack = pd.concat([sep_data, good_data], axis=0)
# Place the DataFrames side by side
horizontal_stack = pd.concat([sep_data, good_data], axis=1)
Row Index Values and Concat
Have a look at the vertical_stack dataframe? Notice anything unusual?
The row indexes for the two data frames sep_head and sep_head_last10
have been repeated. We can reindex the new dataframe using the reset_index() method.
Writing Out Data to CSV
We can use the to_csv command to do export a DataFrame in CSV format. Note that the code
below will by default save the data into the current working directory. We can
save it to a different folder by adding the foldername and a slash to the file
vertical_stack.to_csv('foldername/out.tsv'). We use the ‘index=False’ so that
pandas doesn’t include the index number for each line.
# Write DataFrame to CSV
vertical_stack.to_csv('out.tsv', sep='\t', index=False)
Check out your working directory to make sure the CSV wrote out properly, and that you can open it! If you want, try to bring it back into python to make sure it imports properly.
# For kicks read our output back into python and make sure all looks good
new_output = pd.read_csv('out.csv', delimiter='\t')
Joining DataFrames
When we concatenated our DataFrames we simply added them to each other - stacking them either vertically or side by side. Another way to combine DataFrames is to use columns in each dataset that contain common values (a common unique id). Combining DataFrames using a common field is called “joining”. The columns containing the common values are called “join key(s)”. Joining DataFrames in this way is often useful when one DataFrame is a “lookup table” containing additional data that we want to include in the other.
NOTE: This process of joining tables is similar to what we do with tables in an SQL database.
What we will do now is look at an example of joining together two data frames based on an overlapping column. Our two dataframes will be our vertically stacked output, and a second data frame with organism information.
Joining Two DataFrames
To better understand joins, let’s grab the first 10 lines of our data as a
subset to work with. We’ll use the .head method to do this. We’ll also read
in a subset of the sep_data table.
# Read in first 10 lines of sep_data table
sep_head = sep_data.head(10)
# Make a small subset of the good_data frame
organism_info = pd.read_csv('organism_data.tsv', delimiter='\t')
subset = organism_info.head(10)
In this example, sep_data is the lookup table containing the master list of columns.
Identifying join keys
To identify appropriate join keys we first need to know which field(s) are shared between the files (DataFrames). We might inspect both DataFrames to identify these columns. If we are lucky, both DataFrames will have columns with the same name that also contain the same data. If we are less lucky, we need to identify a (differently-named) column in each DataFrame that contains the same information.
>>> sep_head.columns
Index(['Unnamed: 0', 'length', 'sequence', 'quals', 'readName', 'runID',
'readNum', 'channel', 'lane', 'date', 'hours'],
dtype='object')
>>> subset.columns
Index(['Unnamed: 0','readName', 'runID',
'organism'], dtype='object')
In our example, the join key is the column containing the readName which is called readName.
Now that we know the fields with the common species ID attributes in each DataFrame, we are almost ready to join our data. However, since there are different types of joins, we also need to decide which type of join makes sense for our analysis.
Inner joins
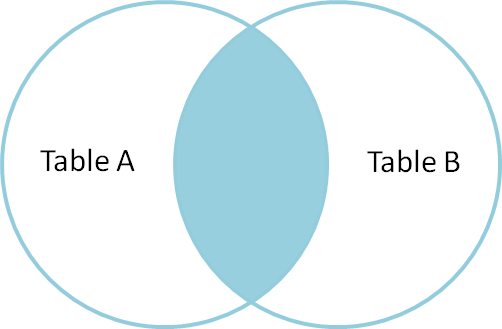
The most common type of join is called an inner join. An inner join combines two DataFrames based on a join key and returns a new DataFrame that contains only those rows that have matching values in both of the original DataFrames.
Inner joins yield a DataFrame that contains only rows where the value being joins exists in BOTH tables. An example of an inner join, adapted from this page is below:
The pandas function for performing joins is called merge and an Inner join is
the default option:
merged_inner = pd.merge(left=sep_head,right=subset, left_on='readNum', right_on='readNum')
# If the same column had different names in the two dataframes,
#we could specifiy this via "left_on" and "right_on"
# What's the size of the output data?
merged_inner.shape
merged_inner
OUTPUT:
Unnamed: 0_x length sequence \
0 0 368 TTATTGTAGTCGGTGGTGTGGCGGGTTGACTGAACTTGCTGCTTTT...
1 1 27977 TTGTTGGCACTTCGTTTCAGTTCTGCGGTGCTGGGCGGCGACCTCG...
2 2 3040 GGATTTCAGTTGCATGTTACTTATCCAAATTGTGTTTGGTTAGTCG...
3 3 13805 ATGCGTACTTCGTTCATTGTACTTCGTTCCAGTTACGTATTGCTGT...
4 5 2731 ATCACCGTTCGAGAGATTACGTATTGCGGATAGCGCCCGTAACCTG...
5 6 3224 CTCATAGGTTTCGTTCGGTTACGGTATTGCTGCCATCAGATTGTGT...
6 7 652 TCGGTACTTCGCGGTTTCGCAGTTACGTATTGCTTGCGTGTGGAAA...
7 8 1216 TTGCGTGGTGAATTCATTCTCCTCGTGAATATCGACTTCAGGACGA...
8 9 10827 TTTTTTTGTGTATTGGGCGGCAGCCTCTTTCGCTATTTATGAAATT...
quals \
0 )""'$#$#&&'+)-&,%%%*&%*+(,*%%)*,-'+*+,.//*($%$...
1 $&%'*'&$%'&(#(+410&+*,$#$$%')()219622522369343...
2 +-&7.)*3537&+%+$4'%%%$(*5)+)*-,*.,,.*'**1.19+'...
3 %*&'))*,204&'''#'''*'),+,*,11&%(+7+2.0,3-*)*'(...
4 %$&*)6:0-+*)+$-'%(**.10'%)'%&''+*16-.76716,)(*...
5 ##((+%'$&&0+.103)3//&,'$%%.'*+233++69+.&4**-/4...
6 ##&#*&*+.+'&+,*,,&%&%,0/./88,8,)****)**,''&))*...
7 )"%%&$'%#%&%++--4;0.,&'&&)&%&$$%&$)'&'%$&.&-$%...
8 $$$$&('(*(-&&('*'323,),,)&''+.+%'))0++''%.--*0...
readName \
0 @bf327144-9003-4b2d-bad5-4cb380f40e8d
1 @f54a0ac6-3563-432f-a447-60e4c95efb79
2 @43a565b2-0dcb-4ea6-a32d-3984c3746e47
3 @f2f91bb1-2592-4193-b9ac-8d94b0f740b1
4 @26ad1236-11c3-40c7-9a6e-13899b65223a
5 @d666315c-d4e0-4f04-b34c-475670d82571
6 @0d00671e-d5db-4c2f-acba-7fe9061448f0
7 @f55e9dec-43f9-45c4-a79f-b04921bbcd84
8 @0bb5a3d8-29c6-4513-b6f5-8ad8a76549e6
runID readNum channel \
0 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=14750 ch=280
1 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=7291 ch=186
2 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=21764 ch=199
3 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=28414 ch=153
4 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=7381 ch=330
5 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=42346 ch=280
6 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=26774 ch=180
7 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=59335 ch=186
8 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=13975 ch=199
lane date hours \
0 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 23:12:05
1 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 21:36:08
2 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 02:20:52
3 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 23:56:01
4 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 21:58:53
5 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 01:29:03
6 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 22:38:44
7 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 02:42:24
8 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 01:38:34
Unnamed: 0_y animal
0 0 Nerodia_fasciata_museum_prep
1 1 Nerodia_fasciata_museum_prep
2 2 Nerodia_fasciata_museum_prep
3 3 Nerodia_fasciata_museum_prep
4 5 Nerodia_fasciata_museum_prep
5 6 Nerodia_fasciata_museum_prep
6 7 Nerodia_fasciata_museum_prep
7 8 Nerodia_fasciata_museum_prep
8 9 Nerodia_fasciata_museum_prep
The result of an inner join of sep_head and subset is a new DataFrame
that contains the combined set of columns from sep_head and subset. It
only contains rows that have two-letter species codes that are the same in
both the sep_head and subset DataFrames. In other words, if a row in
sep_head has a value of readNum that does not appear in the readName
column of species, it will not be included in the DataFrame returned by an
inner join. Similarly, if a row in subset has a value of readName
that does not appear in the readNum column of sep_head, that row will not
be included in the DataFrame returned by an inner join.
The two DataFrames that we want to join are passed to the merge function using
the left and right argument. The left_on='readNum' argument tells merge
to use the readName column as the join key from sep_head (the left
DataFrame). Similarly , the right_on='readName' argument tells merge to
use the readName column as the join key from subset (the right
DataFrame). For inner joins, the order of the left and right arguments does
not matter.
The result merged_inner DataFrame contains all of the columns from sep_head
(record id, month, day, etc.) as well as all the columns from subset
(readName, genus, species, and taxa).
Notice that merged_inner has fewer rows than sep_head. This is an
indication that there were rows in sep_data with value(s) for readName that
do not exist as value(s) for readName in organism_data.
Left joins
What if we want to add information from subset to sep_head without
losing any of the information from sep_head? In this case, we use a different
type of join called a “left outer join”, or a “left join”.
Like an inner join, a left join uses join keys to combine two DataFrames. Unlike
an inner join, a left join will return all of the rows from the left
DataFrame, even those rows whose join key(s) do not have values in the right
DataFrame. Rows in the left DataFrame that are missing values for the join
key(s) in the right DataFrame will simply have null (i.e., NaN or None) values
for those columns in the resulting joined DataFrame.
Note: a left join will still discard rows from the right DataFrame that do not
have values for the join key(s) in the left DataFrame.
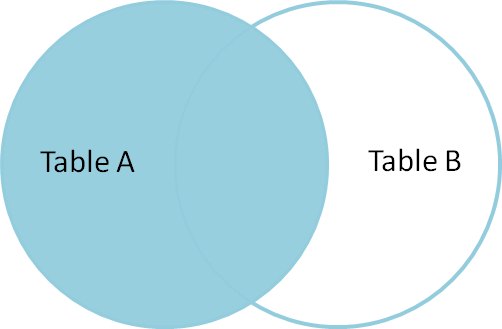
A left join is performed in pandas by calling the same merge function used for
inner join, but using the how='left' argument:
merged_left = pd.merge(left=sep_head,right=subset, how='left', left_on='readNum', right_on='readNum')
merged_left
**OUTPUT:**
Unnamed: 0_x length sequence \
0 0 368 TTATTGTAGTCGGTGGTGTGGCGGGTTGACTGAACTTGCTGCTTTT...
1 1 27977 TTGTTGGCACTTCGTTTCAGTTCTGCGGTGCTGGGCGGCGACCTCG...
2 2 3040 GGATTTCAGTTGCATGTTACTTATCCAAATTGTGTTTGGTTAGTCG...
3 3 13805 ATGCGTACTTCGTTCATTGTACTTCGTTCCAGTTACGTATTGCTGT...
4 4 17176 TTGGTACAGCCACTTCGTTCAGTTACGTATTGCTGGCGGCGACCTC...
5 5 2731 ATCACCGTTCGAGAGATTACGTATTGCGGATAGCGCCCGTAACCTG...
6 6 3224 CTCATAGGTTTCGTTCGGTTACGGTATTGCTGCCATCAGATTGTGT...
7 7 652 TCGGTACTTCGCGGTTTCGCAGTTACGTATTGCTTGCGTGTGGAAA...
8 8 1216 TTGCGTGGTGAATTCATTCTCCTCGTGAATATCGACTTCAGGACGA...
9 9 10827 TTTTTTTGTGTATTGGGCGGCAGCCTCTTTCGCTATTTATGAAATT...
quals \
0 )""'$#$#&&'+)-&,%%%*&%*+(,*%%)*,-'+*+,.//*($%$...
1 $&%'*'&$%'&(#(+410&+*,$#$$%')()219622522369343...
2 +-&7.)*3537&+%+$4'%%%$(*5)+)*-,*.,,.*'**1.19+'...
3 %*&'))*,204&'''#'''*'),+,*,11&%(+7+2.0,3-*)*'(...
4 $$*'+'&&'&%&).412183012;1118/9//.;6;6587466:2+...
5 %$&*)6:0-+*)+$-'%(**.10'%)'%&''+*16-.76716,)(*...
6 ##((+%'$&&0+.103)3//&,'$%%.'*+233++69+.&4**-/4...
7 ##&#*&*+.+'&+,*,,&%&%,0/./88,8,)****)**,''&))*...
8 )"%%&$'%#%&%++--4;0.,&'&&)&%&$$%&$)'&'%$&.&-$%...
9 $$$$&('(*(-&&('*'323,),,)&''+.+%'))0++''%.--*0...
readName \
0 @bf327144-9003-4b2d-bad5-4cb380f40e8d
1 @f54a0ac6-3563-432f-a447-60e4c95efb79
2 @43a565b2-0dcb-4ea6-a32d-3984c3746e47
3 @f2f91bb1-2592-4193-b9ac-8d94b0f740b1
4 @040557c3-1df7-475b-84e5-4ce2ea532508
5 @26ad1236-11c3-40c7-9a6e-13899b65223a
6 @d666315c-d4e0-4f04-b34c-475670d82571
7 @0d00671e-d5db-4c2f-acba-7fe9061448f0
8 @f55e9dec-43f9-45c4-a79f-b04921bbcd84
9 @0bb5a3d8-29c6-4513-b6f5-8ad8a76549e6
runID readNum channel \
0 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=14750 ch=280
1 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=7291 ch=186
2 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=21764 ch=199
3 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=28414 ch=153
4 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=48784 ch=153
5 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=7381 ch=330
6 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=42346 ch=280
7 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=26774 ch=180
8 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=59335 ch=186
9 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=13975 ch=199
lane date hours \
0 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 23:12:05
1 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 21:36:08
2 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 02:20:52
3 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 23:56:01
4 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 02:11:59
5 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 21:58:53
6 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 01:29:03
7 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-16 22:38:44
8 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 02:42:24
9 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 01:38:34
Unnamed: 0_y animal
0 0 Nerodia_fasciata_museum_prep
1 1 Nerodia_fasciata_museum_prep
2 2 Nerodia_fasciata_museum_prep
3 3 Nerodia_fasciata_museum_prep
4 NaN NaN
5 5 Nerodia_fasciata_museum_prep
6 6 Nerodia_fasciata_museum_prep
7 7 Nerodia_fasciata_museum_prep
8 8 Nerodia_fasciata_museum_prep
9 9 Nerodia_fasciata_museum_prep
The result DataFrame from a left join (merged_left) looks very much like the
result DataFrame from an inner join (merged_inner) in terms of the columns it
contains. However, unlike merged_inner, merged_left contains the same
number of rows as the original sep_head DataFrame. When we inspect
merged_left, we find there are rows where the information that should have
come from subset (i.e., readNum) is
missing (they contain NaN values):
merged_left[ pd.isnull(merged_left.animal) ]
**OUTPUT:**
Unnamed: 0_x length sequence \
4 4 17176 TTGGTACAGCCACTTCGTTCAGTTACGTATTGCTGGCGGCGACCTC...
quals \
4 $$*'+'&&'&%&).412183012;1118/9//.;6;6587466:2+...
readName \
4 @040557c3-1df7-475b-84e5-4ce2ea532508
runID readNum channel \
4 runid=0adce96f0fe4a9964393668c94df10a3f88b9d25 read=48784 ch=153
lane date hours \
4 0//20180216_FAH50339_MN24138_sequencing_run_la... 2018-02-17 02:11:59
Unnamed: 0_y animal
4 NaN NaN
These rows are the ones where the value of animal from sep_head (in this
case, PF) does not occur in subset.
Other join types
The pandas merge function supports two other join types:
- Right (outer) join: Invoked by passing
how='right'as an argument. Similar to a left join, except all rows from therightDataFrame are kept, while rows from theleftDataFrame without matching join key(s) values are discarded. - Full (outer) join: Invoked by passing
how='outer'as an argument. This join type returns the all pairwise combinations of rows from both DataFrames; i.e., the result DataFrame willNaNwhere data is missing in one of the dataframes. This join type is very rarely used.
Key Points